A big data toolset for large-scale biomedical text mining
1. IntroductionA large number of biomedical research articles are published every day, accumulating rich information, such as genetic variants, genes, phenotypes, diseases, and treatments. Rapid yet accurate text mining on large-scale scientific literature can discover novel knowledge to better understand human diseases, and to improve the quality of disease diagnosis, prevention, and treatment. We designed and developed an efficient text mining framework called SparkText on a Big Data infrastructure, which is composed of Apache Spark data streaming and machine learning algorithms, combined with Apache Cassandra NoSQL database. SparkText is designed for mining large-scale scientific articles published on multiple journals. To demonstrate its performance for classifying cancer types, we extracted information (e.g., breast, prostate, and lung cancers) from thousands of articles, and then employed three different classification algorithms, including Naive Bayes, Support Vector Machine (SVM), and Logistic Regression, to build prediction models using 5-fold cross validation on 29,437 full text articles downloaded from the PubMed. The accuracy of predicting a cancer type by SVM using the 29,437 full text articles was 93.81%. Compared to other tools that took more than 11 hours to mine the 29,437 full text articles, SparkText took about 6 minutes to mine the same dataset. It demonstrated the potential of mining large-scale scientific articles on a Big Data infrastructure, with real-time update by new articles that are published every day. SparkText can be extended to other areas of biomedical research. SparkText is developed by Dr. Max M. He et al.. This toolset is free to academic community. Please feel free to contact Max with any questions, feedback, or bug reports at max.he at m2kgenomics.com We are interested to hear your comments and feedback as features and improvements will be added in future releases. If you have ideas for improving SparkText or features you wish to add, we will be glad to hear about them. 2. Highlights in Biomedical Text Mining
3. SparkText Architectural DesignA brief architectural design of the SparkText is shown in the following: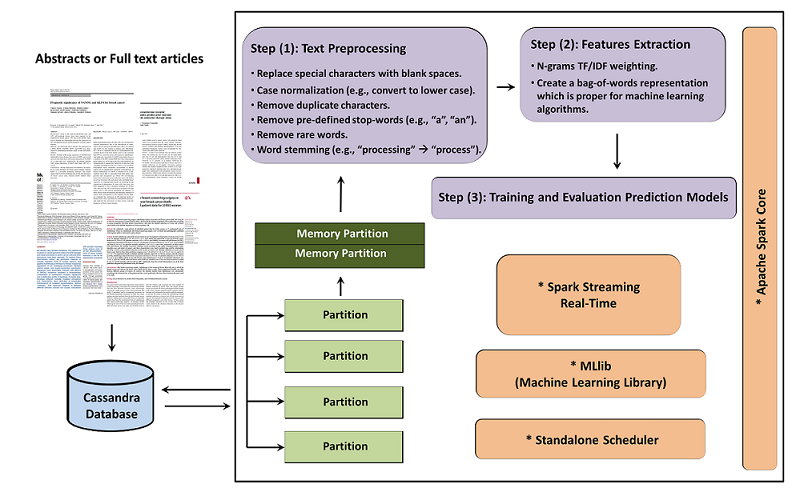
4. InstallationSparkText is developed using Java SE 8 as an Apache Maven project. In case you want to work with the source code which is available here, you need at least one Java programming IDE (e.g., Eclipse, NetBeans) and JDK 8. Otherwise, and in case you want to run the executable JAR file, you need JRE 8. As SparkText builds on top of Apache Spark and Apache Cassandra database, which itself relies on Spark for job execution, the installation/usage definitely requires a working Spark and Cassandra database. To install Cassandra database, please refer to here. SparkText will automatically create a Cassandra keyspace (DB) namely "sparktext". 4.1. DependenciesAll dependencies embedded in pom.xml file as follows:5. DownloadSparkText-1.0.0 is the latest version. Users can download the JAR file here. Source code of SparkText can be downloaded after obtaining a license agreement with Marshfield Clinic Applied Sciences. To run the executable JAR file through the command line, you need to use the following command: 6. User ManualA user manual is also available here as a PDF file.7. A Sample DatasetA sample dataset that includes 19,681 abstracts is available here while another sample dataset that includes 29,437 full text articles is available here as a .zip file. The dataset stored as a single .csv file. The entire content of the dataset downloaded from the FTP Service of PubMed Central (PMC). 8. Terms of UsageSparkText is freely available for any academic, research, and educational purposes. Terms of usage include: 1) You agree to cite the following paper if the package is used for published research. 2) You agree to use the SparkText package only for academic, research, and educational purposes. 3) You agree to not distribute the package. 4) You agree that the code may not be modified or used for non-academic purposes without prior approval.9. CitationsYe Z, Tafti AP, He KY, Wang K, He MM*. Sparktext: Biomedical text mining on big data framework. PLoS One, 2016 11(9), e0162721 [View PubMed] 10. Useful Links |